Superenhancers in Cancer Therapy Resistance
Identification of CRE Regulatory Elements Driving Cancer Resistance to Treatment
Published: 12 September 2025
RESEARCH PAPER (HOPEFULLY) COMING SOON …
In recent years, enhancer reprogramming has been identified as a driver of carcinogenesis. Multiple reseach papers showed that abberrant transciptional programs are required in malignant cells, and that their targeting presents a promising therapeutic strategy. Nonetheless, simply choosing “the ideal” enhancer for a therapeutic purpose is not that simple, due to the complexity of enhancer networks and the related redundancy, their dynamic nature and the enhancer reprogramming.
Super-Enhancers and the Puzzle of Cancer Therapy Resistance
Cancer treatments often stumble when tumours adapt and evade attack. Recent studies highlight super-enhancers (SE) as a key factor in this resistance.
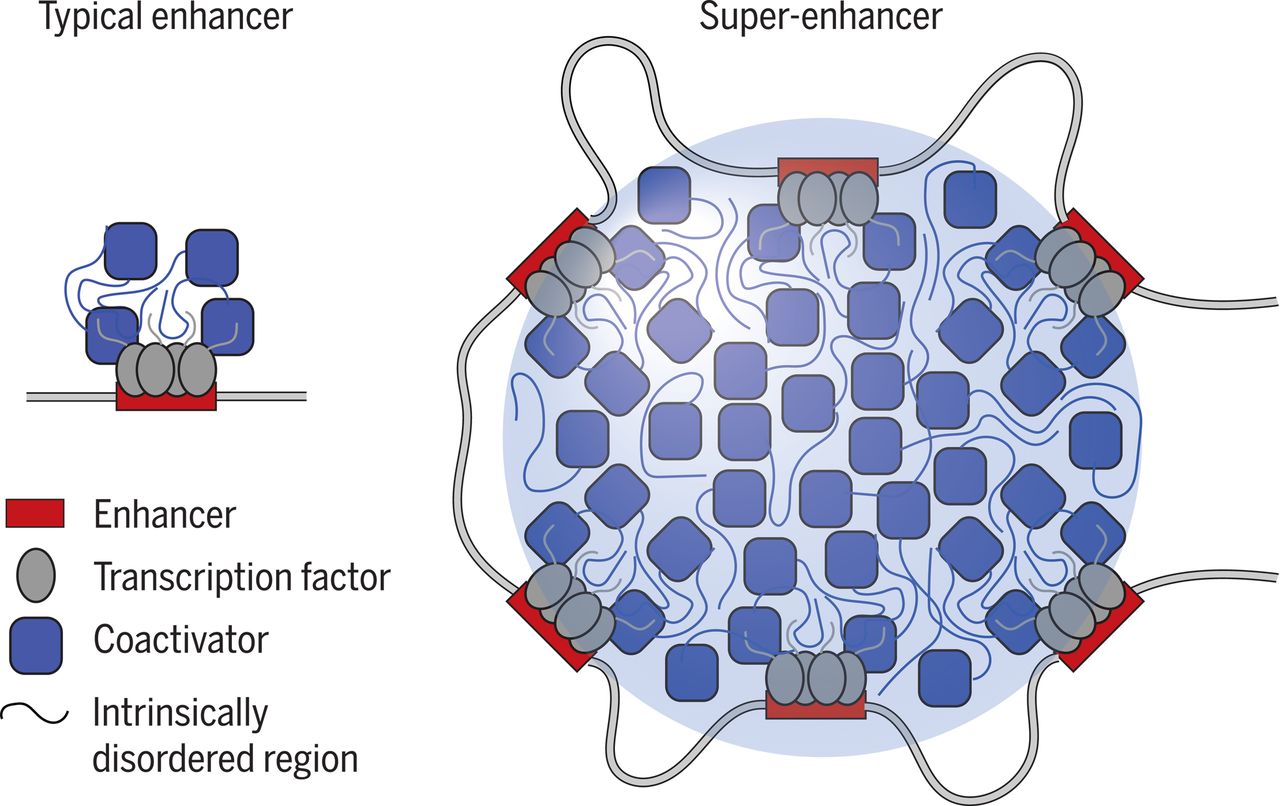
SEs are dense clusters of regulatory DNA that recruit unusually high numbers of transcription factors and co-activators. This concentrated activity drives the powerful expression of genes, including those that keep cancer cells alive. When therapy applies pressure, these elements help tumours rewire their transcriptional networks to survive.
Several mechanisms underlie this escape. Chromosomal rearrangements can reposition SEs next to oncogenes, intensifying their output. Tiny mutations may create new enhancer sites that attract transcription factors, switching on resistance-related genes. Cancer cells can also assemble alternative transcriptional complexes, enabling gene activation even when drugs target BRD4 or CDK7, proteins crucial for SEs function.
Targeted inhibitors such as JQ1 and THZ1 aim to disrupt these powerful regulatory hubs and have shown encouraging effects in pre-clinical models. Yet tumours frequently evolve counter-measures, prompting interest in combined approaches, for example pairing BRD4 inhibitors with PARP1 or immune-checkpoint therapies.
SEs therefore represent both a challenge and an opportunity. Understanding how they maintain malignant gene programmes offers a route to more durable treatments and a clearer strategy for overcoming resistance that has long undermined cancer therapy.
This project aimed to answer a number of questions:
- What deep learning frameworks can be utilised to capture the SE and enhancer sites which impact the tumour progression and affect treatment?
- Are there any particularily powerful SEs which are only present in a tumour subpopulation which drives the cancer progression/metastasis? Can we identify those?
- Can introduce novel treatment strategies or repurpose drugs to demonstrate synergistic effects in preclinical models? How would that improve patient’s prognosis?
The answers to that question resulted in drafting a research paper and subbmitting it to a peer reviewed journal.