Personalised Medicine Approaches in Ovarian Cancer
with a focus on HGSOC
Published: 1 September 2025
High-grade serous ovarian cancer (HGSOC) remains one of the most lethal gynaecologic malignancies, with poor survival rates and few effective treatments. Rather than being driven by a high mutational load, HGSOC progression is often governed by epigenetic dysregulation and enhancer reprogramming, processes that foster tumour plasticity and therapeutic resistance. These enhancer and super-enhancer elements represent critical levers for tailoring treatment to an individual patient’s tumour biology.
One of the promises in treatment of HGSOC is personalised medicine.
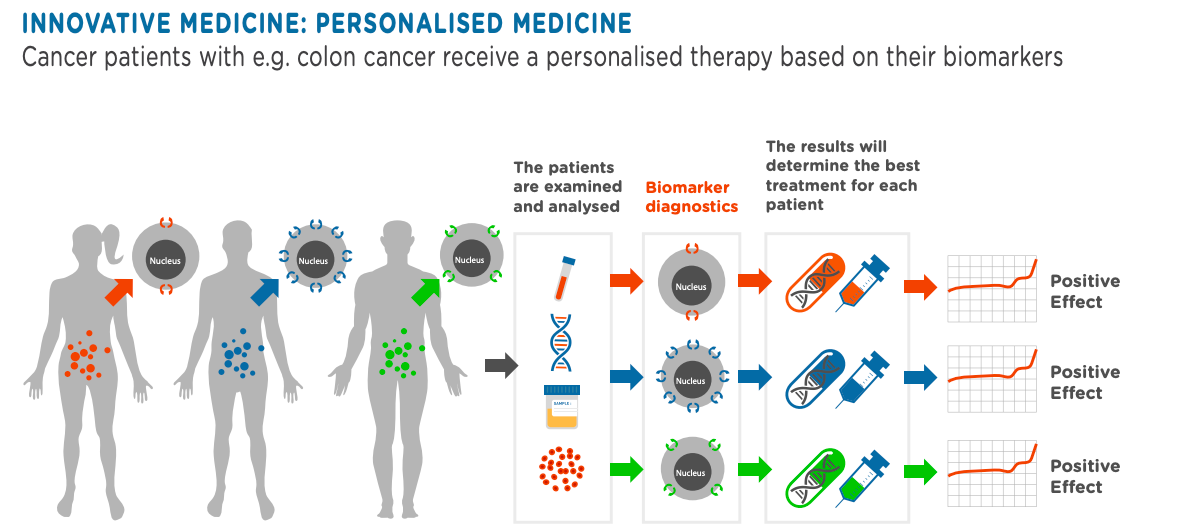
Personalised medicine focuses on matching the most suitable therapy and dose to the needs of each individual patient. Rather than adopting a “one-size-fits-all” strategy, it recognises that people vary in their genes, environment and lifestyle, and that these differences influence how they experience illness and respond to treatment.
While the terms precision medicine and personalised medicine are sometimes used interchangeably, there is a subtle distinction. Precision medicine concentrates on identifying the molecular or genetic features of a disease in order to select targeted interventions, often at the level of a patient subgroup. Personalised medicine, in contrast, goes a step further by considering the whole person, including their preferences, social context and overall health, to design a plan that is both scientifically targeted and individually appropriate.
Beyond choosing therapies, personalised medicine also aims to sharpen diagnostic accuracy, estimate an individual’s risk of developing a condition, and intervene early to prevent progression or complications.
This work outlines machine-learning–enabled frameworks for personalised medicine, designed to single-cell datasets in order to expose patient-specific therapeutic vulnerabilities. Multi-omic data, including single-cell epigenomic and transcriptomic profiles, were integrated to map non-coding somatic single nucleotide variants (SNVs) in cis-regulatory elements (CREs) and their interactions at cellular resolution. Dimensionality reduction and lineage tracking methods revealed malignant subpopulations that may demand distinct therapeutic strategies.
To capture CREs-driven dependencies, deep neural networks (convolutional and recurrent architectures) were trained for sequence-based CREs classification, with attention layers highlighting key transcription factors in each patient sample. Transfer learning from large reference epigenomic resources (ENCODE/ROADMAP) allowed accurate modelling even in data-scarce ovarian cohorts, a crucial step for real-world personalised care.
Downstream integration with transcription-factor scoring, DepMap essentiality screens, and protein–interaction networks (STRING, CORUM, STITCH) pinpointed patient-specific CREs-regulated transcription-factor complexes and druggable pathways.
Together, these enhancer-centric, transfer-learning ML approaches demonstrate how regulatory-genomic insights can be translated into precision treatment plans, bringing truly personalised medicine closer to reality for individuals with high-grade serous ovarian cancer.